The cst cell Array
The constraints of all defined volumes of interest (VOIs) are stored inside the cst cell array. It is structured as follows:
Screenshot of the cst-cell:
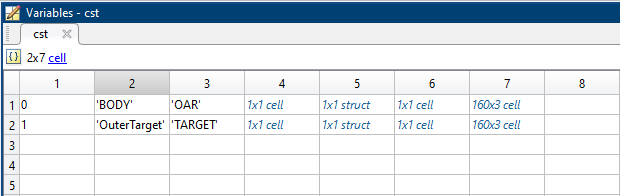
Column |
Content |
Description |
|---|---|---|
1 |
Number to identify the VOI |
|
2 |
String describing the VOI |
|
3 |
Specification whether the VOI is an organ at risk (OAR), a target volume or should be ignored |
|
4 |
Vectors containing the indices of all voxels of the CT that are covered by the VOI. Stored as a cell array of vectors (for enabling handling of multiple scenarios) |
|
5 |
Structure containing information about the tissue of the VOI and its overlap priority |
|
6 |
Cell array containing information about the functions used to calculate the objective & constraint function value |
|
7 |
Precomputed Contours |
After GUI startup, this column contains precomputed contour data for display |
VOI index
All defined VOIs are enumerated starting with 0.
VOI name
The VOI name is a string containing an organ name or a short description of the volume (e.g. BODY, Liver, GTV, …).
VOI type
The VOI type specifies how the volume is considered during treatment planning:
VOI type |
Handling during treatment planning |
|---|---|
TARGET |
The VOI will be covered with spot positions (protons / carbon ions) and bixels (photons) as defined in the stf struct. During the fluence optimization, it will be considered according to the defined dose objectives. |
OAR |
The VOI will not be covered with spot positions or bixels. During the fluence optimization, it will be considered according to the defined dose objectives. |
IGNORED |
The VOI will not be considered during the treatment planning. |
Voxel indices
The indices of all voxels (of the CT-cube) that are covered by the VOI are stored in a vector within a cell array. I.e. we store the segmentation for the VOI as a binary mask, the polygon contour data is not part of matRad’s standard data sets. As the same voxel can be covered by more than one VOI, an overlap priority (see tissue parameters) is defined to handle potential discrepancies when calculating the objective function value and generating the stf struct.
Tissue parameters
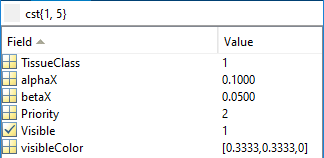
Data can also be stored as in the old format (see below).
New constraints or objectives can be implemented by adding a respective class definition to the matRad.optimization.DoseConstraints or matRad.optimization.DoseObjectives folder.
Dose Objectives & Constraints since v2.10.0
matRad supports inverse planning based on the minimization of a weighted sum of objectives subject to non-linear yet differentiable hard constraints. The following kind of individual objectives are currently supported:
Objective |
Class |
Effect on the objective function |
|---|---|---|
square underdosing |
Only dose values lower than the threshold dose for this VOI are considered for the objective function. The deviations are squared, multiplied with the penalty factor and added to the objective function value. The penalty is normalized to the number of voxels per VOI. |
|
square overdosing |
Only dose values larger than the threshold dose for this VOI are considered for the objective function. The deviations are squared, multiplied with the penalty factor and added to the objective function value. The penalty is normalized to the number of voxels per VOI. |
|
square deviation |
All deviation from a reference dose for this VOI are considered for the objective function. All deviations are squared, multiplied with the penalty factor and added to the objective function value. The penalty is normalized to the number of voxels per VOI. |
|
mean |
All dose values inside this VOI are weighted with the specified, and normalized (see above), penalty factor and added to the objective function value. |
|
EUD |
EUD is the abbreviation for equivalent uniform dose. For this method a weighting factor and an exponent a have to be defined. For the calculation of the objective function value the dose in each voxel is taken to the power of a. Then the sum of all these values is taken (Σ Dia) and divided by the number of voxels. The a-th root of this value is then multiplied with the weighting factor and added to the objective function value. |
|
Min/max DVH objective |
|
Only deviations from the reference dose over/under a reference volume are considered according to Wu & Mohan (2000 Medical Physics). |
Constraints are somewhat built around similar goals as obejctives:
Constraint |
Class |
Description |
|---|---|---|
Min/Max Dose |
Keeps dose above and below the set minimum and maximum dose. Can use a LogSumExp Approximation or a voxel-wise constraint. |
|
Min/Max Mean Dose |
Keeps the mean dose above and below the set minimum and maximum mean dose. |
|
Min/Max EUD |
Keeps the EUD above and below the set minimum and maximum EUD. The EUD is calculated using the same method as for the EUD objective. |
|
min/max DVH |
Keeps the dose volume histogram above and below the set minimum and maximum volunme for a given dose level. |
When generating an objective / constraint from code, we suggest to wrap the instantiation of the objective/constraint in a struct() call, as shown in the first phantom example:
47objective1 = struct(DoseObjectives.matRad_SquaredDeviation(800,45));
48objective2 = struct(DoseObjectives.matRad_SquaredOverdosing(400,0));
49objective3 = struct(DoseObjectives.matRad_SquaredOverdosing(10,0));
This will ensure that, when saving to a mat-file, we don’t save the class object, which improves compatibility.
Before Version 2.10.0
In the earlier version, matRad stored the objectives and constraints defined for inverse planning as an array of structs. matRad_convertOldCstToNewCstObjectives() can be used to convert the old definition to the new format.
Default cst-values
The patient data contained within matRad (ALDERSON, BOXPHANTOM, HEAD_AND_NECK, LIVER, PROSTATE and TG119) have default values defined within the cst-cell.
These values are chosen to produce a reasonable treatment plan, when using coplanar and equidistant photon beams. They can be used as a reference point for more sophisticated treatment plans.